Menu
Menu

The assay leverages in situ DNA organization inside of the nucleus. Extracted DNA does not preserve this organization and therefore not compatible with the assay.
Flash freeze the tissue in liquid nitrogen upon collection and store at -80°C until ready to use.
The workflow has been validated down to 100K cells. No protocol changes are required when using less than 1M cells. What do I give up by using less cells? Less cells means you will recover lower yields after proximity ligation. This ultimately means your libraries may have lower complexity (i.e. higher duplication rate).
No, the LinkPrepTM Kit does not support the use of third-party library preparation kits. The Library Preparation and Primer Modules are part of the core kit offering.
The use of a swinging bucket rotor:
If QC sequencing, the library can be shallow sequenced to 1- 10 million read pairs (2 x 150 bp).
For standard sequencing, we recommend to sequence one library up to 300 million read pairs (2 x 150 bp). The depth depends on the variants you want to detect and its tumor fraction and variant allele frequency.
| Tumor Fraction | Genomic Coverage | # Libraries | # read pairs (2 x 150 bp) | |
| SVs | >50% | 10X | 1 | 100M |
| SVs, SNVs, Indels, CNVs | >50% | 30X | 1 | 300M |
| SVs | 20% – 50% | 30X | 1 | 300M |
| SVs, SNVs, Indels, CNVs | 20% – 50% | 80X | 3 | 800M |
Detection of Low VAF events are dependent on sequence depth
LinkPrepTM Kit users can analyze the data using standard NGS tools. Some recommendations that we have can be found on the following readthedocs. Page: https://varilink.readthedocs.io/en/latest/.
Micro-C uses micrococcal nuclease (MNase) to digest chromatin instead of the restriction enzymes other Hi-C methods use. This approach provides superior signal–to–noise, enabling nucleosome resolution/nucleosome phasing and a more granular view of chromatin folding that includes enhancer-promoter/promoter-promoter interactions and loop extrusion features.
The standard input for Micro-C assay is one million cells.
You will get 8 reactions with each Micro-C kit.
It only takes 1.5 days to go from sample to sequencing-ready library!
The Micro-C workflow does not require sonication. The fragmentation is enzymatic and is carried out at Stage 1 of the protocol.
Since your experimental design will determine the types of downstream analyses available to you, it is critical to consider both in unison prior to generating any data. You can find a detailed discussion of the Micro-C experimental design here.
Micro-C libraries are Illumina compatible. We recommend sequencing 2 x 75 bp, 2 x 100 bp, or 2 x 150 bp.
Dovetail has compiled a comprehensive best practices step-by-step workflow with full documentation that covers:
Dovetail tools for QC and Contact Matrix generation require the Micro-C raw sequence data (*.fq.gz) and a reference genome assembly (*.fa).
Due to the use of restriction enzymes and sonication, traditional Hi-C valid reads require the insert size (or physical coverage) of a read-pair to be greater than the library fragment and to be adjacent to a restriction site. Micro-C libraries do not use restriction enzymes or require sonication (a key source of noise in Hi-C libraries). Therefore, a valid Micro-C read only requires that the insert size (or physical coverage) of a read-pair be greater than the library fragment. Check out this white paper to learn more.
The Dovetail HiChIP MNase assay leverages the Micro-C (MNase-based Hi-C) workflow to conduct chromatin digestions. With the HiChIP MNase assay:
Inclusion of ChIP-seq data simplifies data QC and interpretation but is not an absolute requirement. You can use publicly available data from sources such as ENCODE if you have not generated ChIP-seq data. However, it will not be truly reflective of your experiment or sample of interest.
This will vary depending on the antibody/protein of interest. See table below.
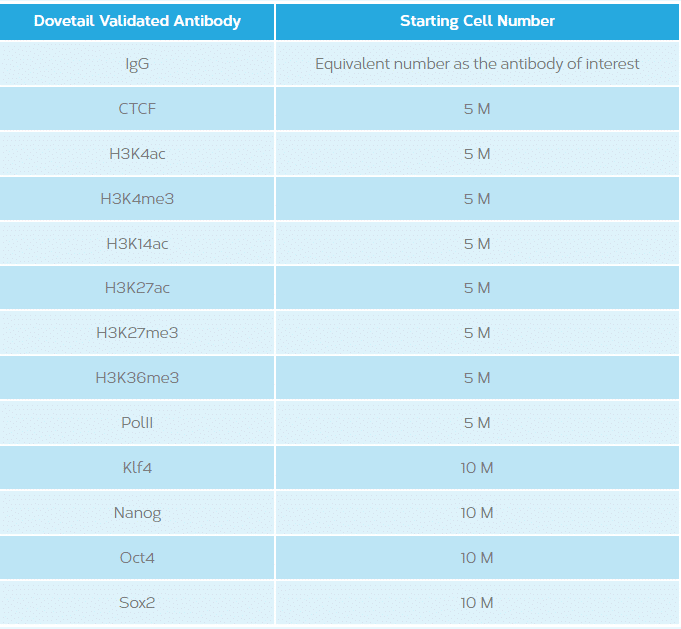
This will vary depending on the occurrence rate of protein-DNA interactions of the protein of interest and the genome in question. The following are recommended starting guidelines:
Dovetail has compiled a comprehensive best practices step-by-step workflow with full documentation that covers:
The Dovetail HiChIP QC tool that assesses the success of a Dovetail HiChIP library requires the raw HiChIP sequence data (*.fq.gz), a reference genome (*.fa) and a file of 1D ChIP-seq peak locations (*.bed).
The standard input for Omni-C is one million cells. A low input protocol is also supported down to one thousand cells.
Cells, animal and plant tissues, and blood.
| Sample | Type | Preparation |
| Cells | Cells | Harvest, wash with 1X PBS, count, aliquot 1M cells, spin down , discard the supernatant and freeze the pellet at -80ºC |
| Animal Tissues | When possible, non-muscle tissues (such as liver, kidney, brain, spleen) are preferred as they generate a higher chromatin yield compared to muscle tissues (such as heart) | Flash freeze in liquid nitrogen immediately upon collection and store at -80ºC until use. |
| Animal Blood | From most to least preferable: Fresh Blood Frozen Blood |
Fresher the better, collect with anticoagulant EDTA, ACDA or Heparin tubes. Fresh blood that cannot be processed within 72 hours of collection, should be frozen. |
| Plant Tissues | From most to least preferable: – Leaves at one or two leaf seedling stage – Very young leaves from more mature plants – Leaves collected from plants that are pretreated in dark for 2-3 days – Tissue from young plant parts other than leaves |
Flash freeze in liquid nitrogen immediately upon collection and store at -80ºC until use. |
| Anthropods | From most to least preferable: – Embryos – Newly hatched larvae – Early pupae – Adults |
Degut or starve adults for 2-3 days prior to freezing. Flash freeze in liquid nitrogen and store at -80ºC until use. |
| Marine Animals/Invertebrates | From most to least preferable: – Internal organs (applicable for larger marine invertebrates) – Newly hatched larvae – Sperm – Blood – Muscle Note: Mantle tissue is not recommended |
Flash freeze in liquid nitrogen and store at -80ºC until use. |
You will get 8 reactions with each Omni-C kit.
The workflow from sample to sequencing–ready library is split over two days.
The Omni-C kit is compatible with hybrid capture approaches. The Omni-C workflow integrates easily with Agilent SureSelect, Illumina TruSight and IDT xGEN.
Omni-C libraries exhibit WGS-like coverage of the genome and do not use restriction enzymes; therefore, you can use off-the-shelf probe sets or design your own probes without an inclusion of a restriction enzyme site.
Yes, frozen cells previously cross-linked with formaldehyde concentration ≥ 1% can be used as input to the assay. Simply skip the formaldehyde fixation step of our recommended two step fixation process.
Dovetail offers library preparation and index primers modules to streamline the library preparation workflow. Omni-C is compatible with third party library preparation kits as well, namely NEBNext® Ultra™ II DNA Library Prep Kit for Illumina®, Kapa® HyperPrep, and Swift Biosciences Accel-NGS® 2S Plus DNA Library Kit.
The Omni-C workflow does not require sonication. The fragmentation is enzymatic and is carried out at Stage 1 of the protocol.
Omni-C libraries are Illumina compatible. We recommend sequencing 2 x 150 bp. One library is typically sufficient for the generation of ~300M total read-pairs. Depending upon final sequencing depth desired, multiple libraries may need to be pooled prior to the sequencing run.
Dovetail has compiled a comprehensive best practices step-by-step workflow with full documentation that covers:
Dovetail tools for QC and Contact Matrix generation require the Omni-C raw sequence data (*.fq.gz) and a draft assembly (genome *.fa).
Due to the use of restriction enzymes and sonication, traditional Hi-C valid reads require the insert size (or physical coverage) of a read-pair to be greater than the library fragment and to be adjacent to a restriction site. Micro-C libraries do not use restriction enzymes or require sonication (a key source of noise in Hi-C libraries). Therefore, a valid Micro-C read only requires that the insert size (or physical coverage) of a read-pair be greater than the library fragment as defined by HiC-Pro. Check out this white paper to learn more.
HiC-Pro is also compatible with Omni-C libraries. Note that there are custom configuration files that are required. Please reach out to us at Dovetail for guidance on how to run HiC-Pro.
This largely depends on application. The following recommendations are good starting points:
Chromosome conformation capture (3C), also known as proximity-ligation, is a molecular biology approach designed to analyze the spatial organization of chromatin. When the ligation events are tagged with biotin and isolated with streptavidin subjected to paired-end whole genome sequencing it is known as Hi-C.
For further information, check out this review paper by Davis et al. (2017).
The Dovetail proximity-ligation kits are user friendly and designed for novice Hi-C users. Plus, our world-class support team is also available to help out if you run into difficulties or have questions.
They are novel Hi-C assays available only from Dovetail Genomics® and differ from traditional Hi-C in the following important ways:
Don’t see your question? Submit it to our support team: [email protected]